Research
Cilia & Ciliopathies
Cilia are microtubule-based organelles that project from nearly every vertebrate cell and mediate critical processes including motility, mechanosensation, and developmental signalling through pathways such as Hedgehog and Wnt. Ciliopathies are a clinically and genetically heterogeneous group of rare diseases caused by structural or functional defects in cilia or basal bodies, encompassing syndromes that affect multiple organ systems simultaneously.
What are ciliopathies?
The Kaplan Lab focuses on a spectrum of ciliopathy syndromes including Joubert syndrome (JBTS), Bardet-Biedl syndrome (BBS), Meckel–Gruber syndrome (MKS), and cranioectodermal dysplasia (CED). These conditions share overlapping genetic architecture — mutations in genes encoding components of the transition zone, IFT machinery, or BBSome — yet produce distinct phenotypes affecting the kidney, retina, brain, and skeleton.
Key discoveries
Approaches & model systems
Why C. elegans?
Caenorhabditis elegans is uniquely suited to ciliopathy research. Its 60 ciliated sensory neurons are anatomically invariant and genetically tractable. The nematode ciliome is highly conserved with humans, CRISPR editing is rapid and inexpensive, and fluorescent reporter strains allow real-time imaging of IFT trains and ciliary membrane proteins in living animals. The lab uses C. elegans to validate variants identified from patient sequencing and to dissect gene function at single-cell resolution.
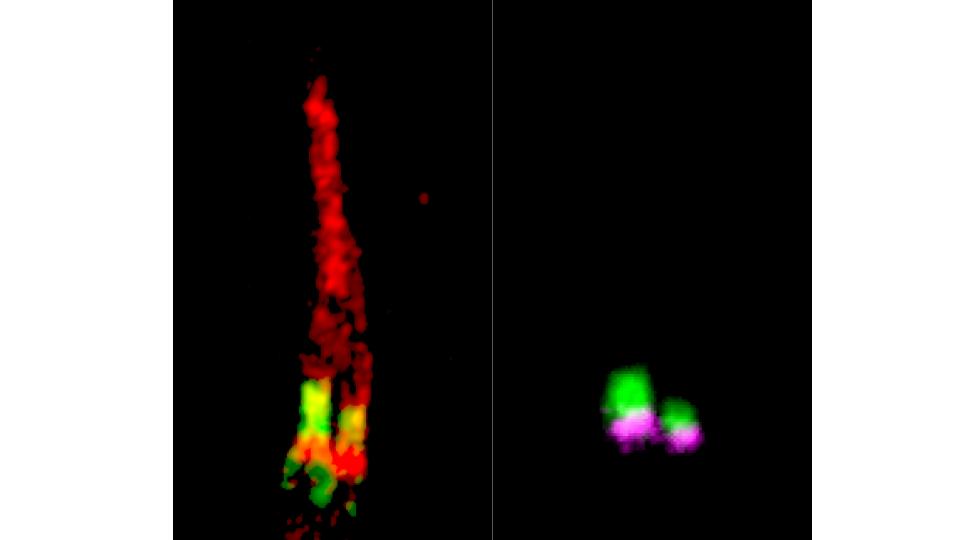
Cilia types in C. elegans sensory neurons.