Research
Disease Modeling
Rare disease diagnosis is hampered by the sheer number of genetic variants of uncertain significance (VUS) discovered by next-generation sequencing. The Kaplan Lab develops and applies a systematic pipeline — combining patient genomics with functional validation in C. elegans — to resolve which variants are truly pathogenic and to understand the mechanisms by which they cause disease.
The variant-to-function pipeline
When a patient presents with a suspected ciliopathy or rare developmental syndrome, whole exome or genome sequencing identifies candidate variants. The challenge is distinguishing causative mutations from benign polymorphisms. The lab's pipeline moves from sequencing to functional answer in three stages:
Candidate variants are filtered by allele frequency, conservation across species, and predicted functional impact. The lab's own ConVarT tool searches for equivalent variants in non-human model organisms to flag those with prior experimental evidence.
The orthologous mutation is precisely introduced into the C. elegans genome using CRISPR/Cas9. The lab uses a PCR-based, cost-efficient screening strategy (see Resources) to confirm correct edits without sequencing every candidate.
Edited worms are assessed for ciliary defects (dye-filling, IFT velocity, cilia morphology by confocal and electron microscopy), developmental phenotypes, and protein localisation using fluorescent reporters. Rescue experiments with wild-type and patient-variant alleles confirm pathogenicity.
Key findings
Technologies used
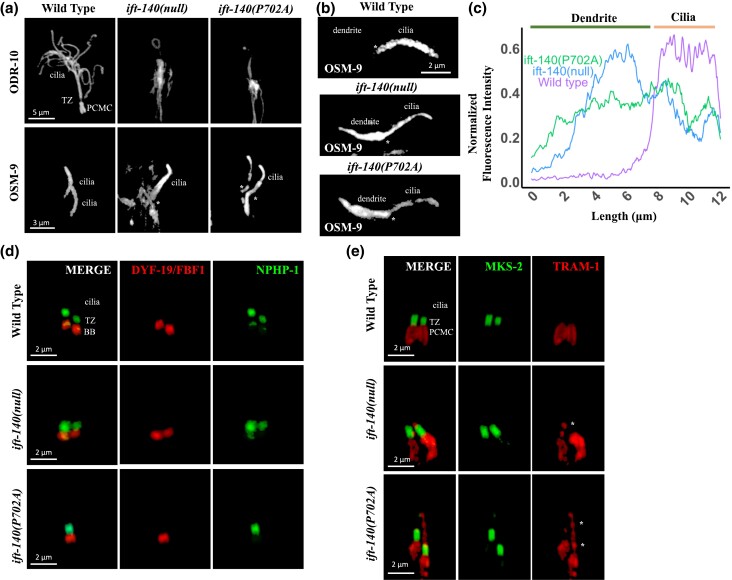
Functional screening of patient variants using C. elegans disease models.